Swimming of micro-organisms
Swimming of micro-organisms is remarkably different from our macroscopic experience. Human swimming overcomes friction of a surrounding fluid by the use of inertia of the fluid and our body. Swimming on a small scale means to overcome friction, where inertia forces are too weak to be of any help. Thus, a micro-organisms swimming strokes are fundamentally different from a macro-organisms’. See the excellent description ‘Life at low Reynolds number’ (E.M. Purcell) and a web search for ‘Swimming at low Reynolds number’ for more information.
A simple self-organized swimmer
Some micro-organisms use cilia and flagella for propulsion. The appendage’s motion is driven by the action of molecular motors. Though motors interact with filaments and elastic elements. Due to the complicated structure of the cellular appendages swimming is insufficiently understood: Somehow the collective action of motor proteins propels micro-organisms. In order to better understand these processes we investigate the dynamics of a simple swimmer as a mathematical model. Three connected spheres compose the swimmer. For a specified motion of the spheres with respect to each other, the swimmer is capable of undergoing directed motion.
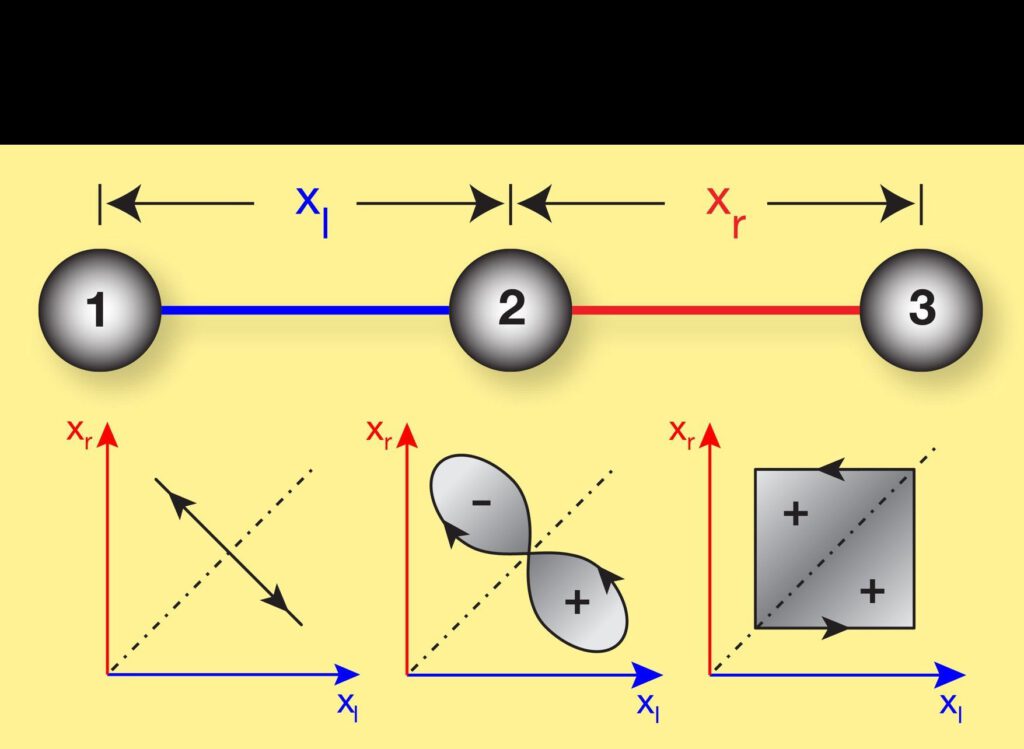
Possible shape changes of the three-sphere swimmer depending on the spheres’ distances left and right. For symmetry reasons the swimmer does not propagate for the motions depicted in the left and middle graph. Only the motion in the right graph allows for directed swimming.
Let the three spheres shall be linked by elements composed of molecular motors, filaments and elastic elements. The motors are coupled elastically together and can bind and unbind from the filaments.
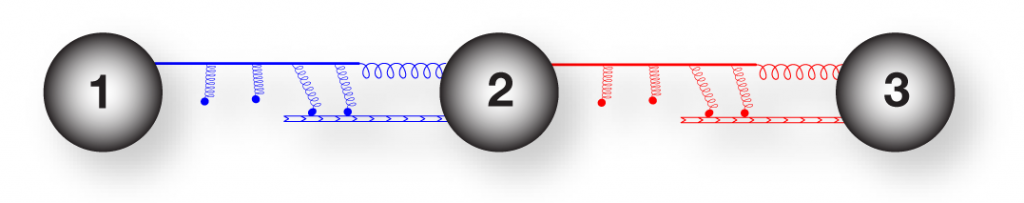
Three sphere swimmer linked by elastic and active motor components.
We find, that the molecular motors can interact in a way so that they induce a motion that is capable of propagating the swimmer. The motion originates from a spontaneous breaking of a dynamic symmetry. Motors self-organize into a state where all motors act together and induce strokes that propagate the swimmer.
References
| Scientific article: A simple self-organized swimmer driven by molecular motors |
| PhD thesis: Self-organized cyclic patterns in muscles and microscopic swimming |